EMIF-METABOLIC
EMIF-Metabolic Project background
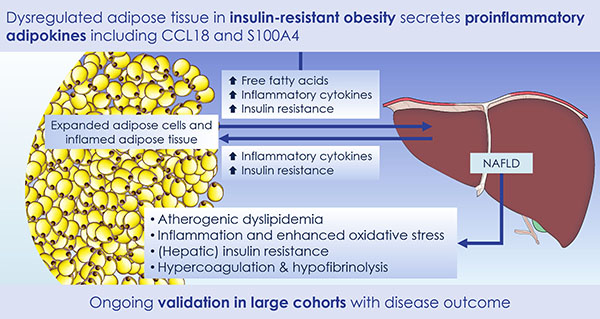
Accumulating evidence indicates that obesity is closely associated with an increased risk of metabolic complications such as type 2 diabetes, coronary heart disease, non-alcoholic fatty liver disease and cancers. However, the association between obesity and any of these complications is complex, with high inter-individual variability in susceptibility to specific metabolic complications of obesity. This association is expected to be additionally influenced by different constitutional, environmental and obesity-specific factors, thus further complicating the development of adequate treatments.
EMIF-Metabolic Project Objectives
 EMIF-Metabolic will focus on two distinct pathways through which individuals who are obese may vary in their risk of the complications of obesity. One is to identify genetic causes of obesity and their relation to the metabolic complications of obesity. The other pathway is to characterize individuals and identify markers associated with metabolic risk irrespective of degree of obesity based on the knowledge that many obese individuals do not become dysmetabolic and insulin resistant. In both instances the complementary approaches of studying extreme phenotypes and population-based cohorts will be followed.
EMIF-Metabolic will focus on two distinct pathways through which individuals who are obese may vary in their risk of the complications of obesity. One is to identify genetic causes of obesity and their relation to the metabolic complications of obesity. The other pathway is to characterize individuals and identify markers associated with metabolic risk irrespective of degree of obesity based on the knowledge that many obese individuals do not become dysmetabolic and insulin resistant. In both instances the complementary approaches of studying extreme phenotypes and population-based cohorts will be followed.
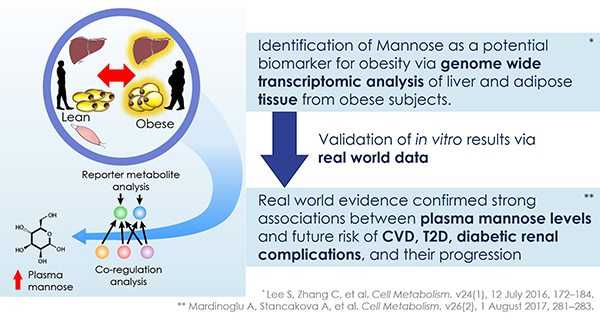
The discovery of predictors of the metabolic complications of adult and paediatric obesity has lead to innovative diagnostic tests, paved the way to novel therapeutics targeted to high-risk individuals, and provided the infrastructure to select individuals for such targeted pharmacological interventions (genetic, epigenetic and ‘omics platforms).
Identified novel mechanisms and potential targets for therapy were characterized in appropriate cell- and animal-based models to facilitate transition to human studies. To this end, EMIF-Metabolic has brought together top European experts in genetics, epidemiology, genomics and other ‘omic’ technologies, human and murine physiology and systems approaches and state-of-the-art technologies.
Work Packages Overview
EMIF-MET
Identify predictors of metabolic complications in obesity
![]()